This page was produced as an assignment for Genetics 564, an undergraduate capstone course at The University of Wisconsin - Madison.
What is a protein domain?
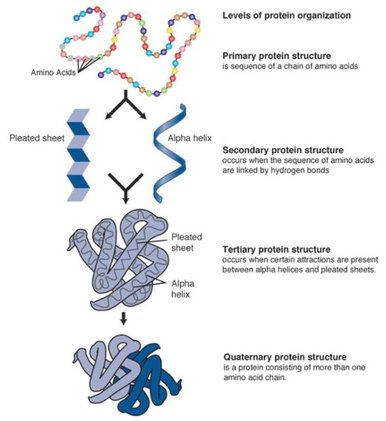
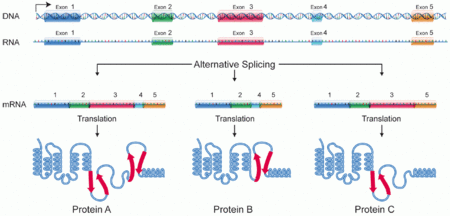
Proteins have three important characteristics that make up their structure. The sequence of amino acids (AA) that are translated from a strand of mRNA make up the protein's primary structure. A protein's secondary structure is makes is determined by the hydrogen bonding between amino acids resulting in alpha helices and beta sheets. The tertiary structure of a protein is the interaction between those alpha helices and beta sheets. A protein domain is part of the protein's tertiary structure, is highly conserved, and plays a major role in its function [1]. Domains have a highly specific and highly conserved three-dimensional structure. Even in different versions of proteins coded from a single gene, domains can have homologies because of similar splicing patterns [2].
How do you study a domain?
There are several databases that identify and categorize protein domains. Two databases (SMART and Pfam) were used to identify the domains present in Amyloid Precursor Protein (APP). Pfam was produced by the European Bioinformatics Institute [3] and uses Markov Models to predict domains from amino acid sequences [4]. A Simple Modular Architecture Research Tool (SMART) is database that specialize in domain structure. It's predictive model generates structures for more than 1200 protein domains [5]. FASTA sequences were obtained for each APP ortholog and were submitted to each database for analysis.
What domains are in APP?
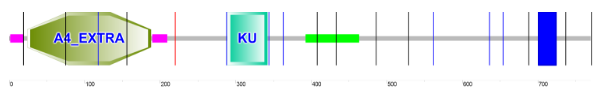
One of the most widely conserved versions of APP is APP 4. It is distinct from other versions because it has one of the longest AA sequences and is widely conserved across a range of model organisms.
How well conserved is the APP4 domain?
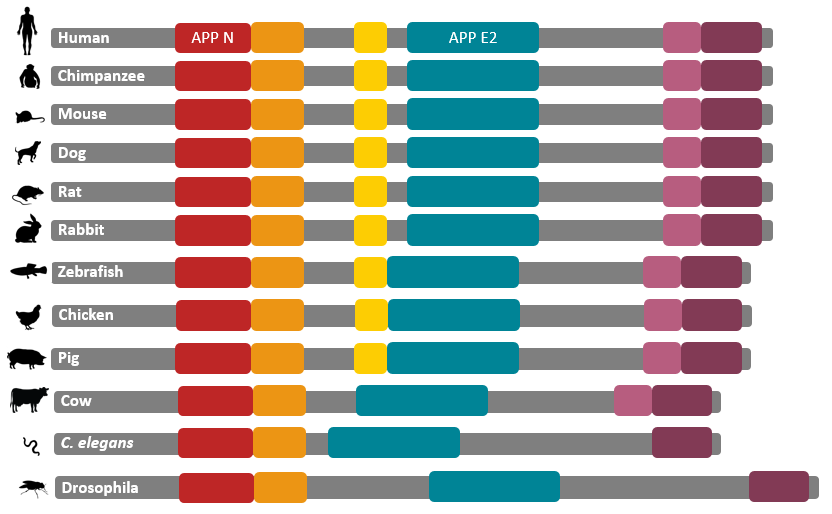
According to the Pfam predictive model, APP N and APP E2 are the most significantly conserved domains. Across the listed model organisms, they are the most well-conserved in terms of spacing and are most significant in terms of size. This diagram helps indicate why Zebrafish embryos are an ideal model organism for further domain analysis experiments. Compared to the Human counterpart, the domains found in Zebrafish are well conserved. Additionally the gene editing in Zebrafish embryos allows for mutagenesis in eukaryotic organisms and the transparent quality of the embryos allows for ideal phenotype viewing in a cost effective manner.
Discussion
For future analysis, investigations involving any of the APP domains should involve either APP N or APP E2 because they are the most widely conserved across the presented model organisms. Therefore confining CRISPR/Cas9 mutagenesis to these regions would be a representative model of mutations in the human counterpart. In addition to using mutagenesis in the two domains to determine their role in APP production, a mutation screen could be used to monitor the interaction of mutant APP with other proteins in the synapse region.
References
Images
Protein Structure: http://www.majordifferences.com/2013/02/difference-between-primary-and.html#.WL95AxLyvR0
Alternative Splicing: https://en.wikipedia.org/wiki/Alternative_splicing
[1] http://www.majordifferences.com/2013/02/difference-between-primary-and.html#.WL-R1hLyvR1
[2] Bork, P. 1991. Shuffled domains in extracellular proteins. Federation of European Biochemical Societies. 286: 1,2
[3] EMBL-EBI. About Pfam. Web. <http://pfam.xfam.org/about>
[4] Truong, K and Ikura, M. 2003. Domain fusion analysis by applying regional algebra to protein sequence and domain databases. BMC Bioinformatics. 4:16.
[5] Letunic, I, Doerks, T, and Bork, P. 2014. SMART: recent updates, new developments, and status in 2015. Nucleic Acids Research. 43, D1: D257-D260.
Protein Structure: http://www.majordifferences.com/2013/02/difference-between-primary-and.html#.WL95AxLyvR0
Alternative Splicing: https://en.wikipedia.org/wiki/Alternative_splicing
[1] http://www.majordifferences.com/2013/02/difference-between-primary-and.html#.WL-R1hLyvR1
[2] Bork, P. 1991. Shuffled domains in extracellular proteins. Federation of European Biochemical Societies. 286: 1,2
[3] EMBL-EBI. About Pfam. Web. <http://pfam.xfam.org/about>
[4] Truong, K and Ikura, M. 2003. Domain fusion analysis by applying regional algebra to protein sequence and domain databases. BMC Bioinformatics. 4:16.
[5] Letunic, I, Doerks, T, and Bork, P. 2014. SMART: recent updates, new developments, and status in 2015. Nucleic Acids Research. 43, D1: D257-D260.