This page was produced as an assignment for Genetics 564, an undergraduate capstone course at The University of Wisconsin - Madison.
What is protein phosphorylation?
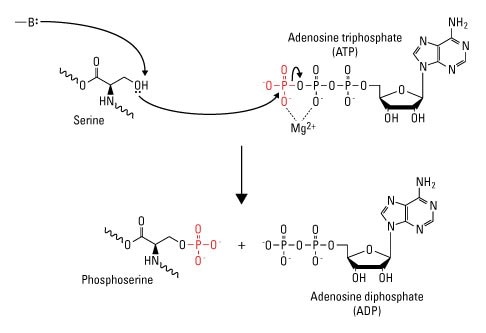
Once a gene is translated into protein, there are a number of chemical modifications that the protein can undergo. These post-translational modifications play a critical role in the cell's regulatory pathways. Phosphorylation plays a critical role in the regulation of many cellular processes including cell cycle, growth, apoptosis and signal transduction pathways, metabolism, division, membrane transport, learning, and memory [1, 2]. Phosphoproteomics is a class of proteomic analysis that identifies and characterizes phosphorylation proteins [1].
|
The reversible process of phosphorylation is specific to three amino acids: serine, threonine, and tyrosine. When phosphorylated they form acid-stable phosphomonoesters [2]. Other amino acid residues like histidine, arginine, and lysine but are less stable and are often undetected by research methods [2]
|
What predicted phosphorylation sites are in APP?
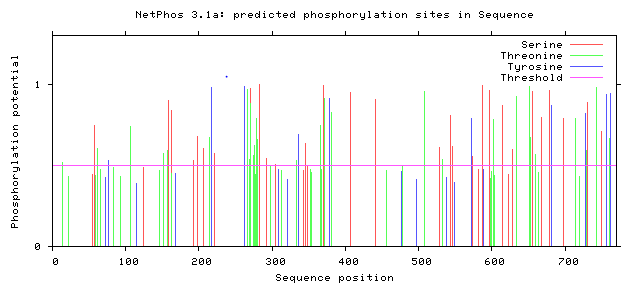
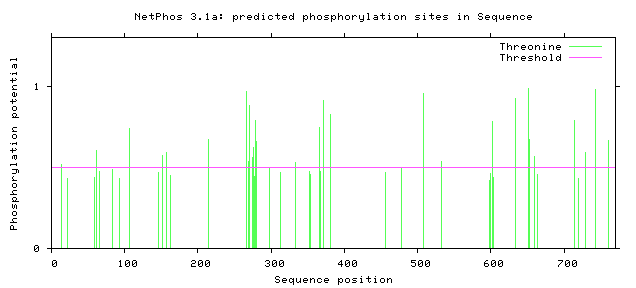
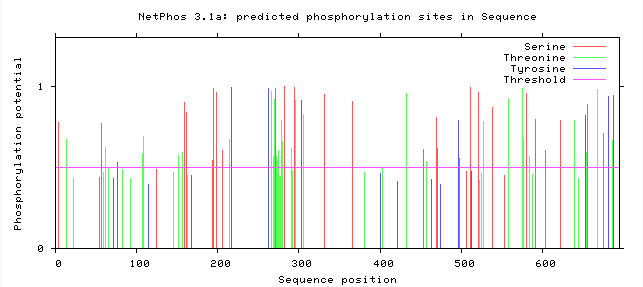
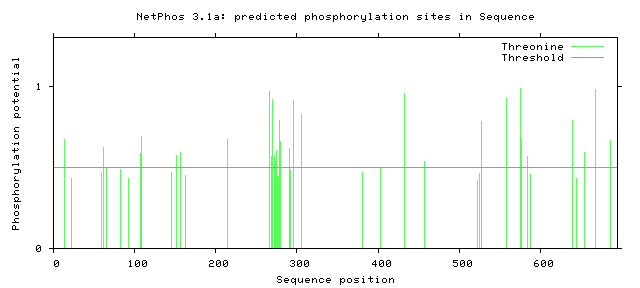
The NetPhos server predicts the phosphorylation sites in eukaryotic proteins specific to three amino acids: Serine, Threonine, and Tyrosine. Knowing these predicted sites can tell us which regions of the protein are subject to enzymatic activity. In the case of the Amyloid Precursor Protein (APP), understanding the phosphorylation sites can help researchers better understand the regulatory and cleavage process that leads to the acceleration of the Amyloid Cascade Hypothesis [3]. Below the predicted phosphorylated regions were gathered in humans and in mice to determine if the predicted residues are consistent in mouse APP.
Discussion
Based on predicted phosphorylation sites using the NetPhos database, we can determine that phosphorylation is consistent across humans and the model organism mice. Although there are slight differences in the predicted phosphorylation sites, this confirms that the use of a mouse model to characterize the regulation of APP with consistent with the human homolog.
References
[1] Thermo Fisher Scientific. 2017. Phosphorylation. Web. https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/phosphorylation.html
[2] Cieśla, J., Frączyk, T., & Rode, W. (2011). Phosphorylation of basic amino acid residues in proteins: important but easily missed. Acta Biochimica Polonica, 58(2), 137-148.
[3] Hardy, J. A., & Higgins, G. A. (1992). Alzheimer's disease: the amyloid cascade hypothesis. Science, 256(5054), 184.
[2] Cieśla, J., Frączyk, T., & Rode, W. (2011). Phosphorylation of basic amino acid residues in proteins: important but easily missed. Acta Biochimica Polonica, 58(2), 137-148.
[3] Hardy, J. A., & Higgins, G. A. (1992). Alzheimer's disease: the amyloid cascade hypothesis. Science, 256(5054), 184.