This page was produced as an assignment for Genetics 564, an undergraduate capstone course at The University of Wisconsin - Madison.
How do you build a protein network?
|
Proteins have a wide range of roles and functions. One of the ways to characterize a protein by it's interaction. Protein - protein interaction varies based on the molecule's biological function and even between normal and disease states. Learning about what our protein of interest - Amyloid Precursor Protein (APP) - interacts with can help us understand it's role across model organisms and in an Alzheimer's disease and normal states. Protein interaction databases like STRING, IntAct, and BioGrid can help you visualize protein interactions that have been testing and are proposed. These networks can be used to determine gaps in your genes function and/or localization by looking for common proteins.
|
How does APP's protein network vary between humans and mice?
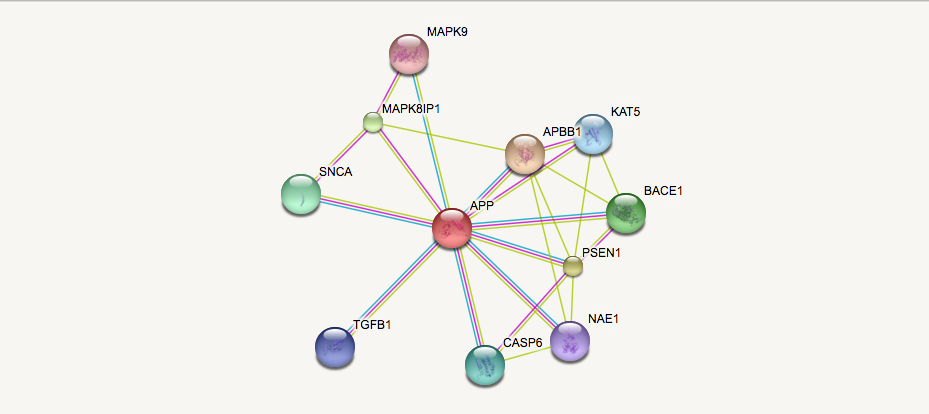
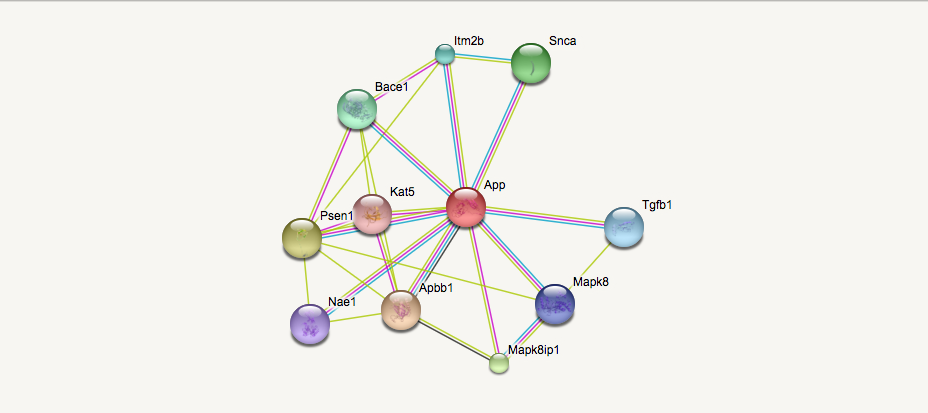
When studying the protein homology of APP between mice and humans we learned that they have a 87.79% identity. However we can further investigate their similarities by comparing and contrasting their protein networks. By analyzing the gene ontology of the proteins within the network, we can gain a better understanding of our primary protein's localization and function.
Discussion
Comparing and contrasting the human and mouse APP STRING Networks revealed two discrepancies. The proteins Itm2b, Mapk8 were found in the mouse network but were not present in the human APP network. However the isoform of Mapk8, Mapk9 is present in the human APP network. UniProt analysis revealed that the protein Itm2b is an integral membrane protein (like APP) that is involved in the processing of the beta-amyloid A4 precursor protein (APP) and acts as an inhibitor of the beta-amyloid peptide aggregation and fibrils deposition. Plays a role in the induction of neurite outgrowth. Functions as a protease inhibitor by blocking access of secretases to APP cleavage sites [1]. In many cases the protein interaction networks of model organisms are better investigated than human protein networks because of the amount of research studies that is performed on them. Further investigation of the protein interaction between APP and Itm2b could reveal new information about the regulation of the Amyloid Cascade Hypothesis.
References
[1] UniProt. 2017. ITM2B Function. [Web]. http://www.uniprot.org/uniprot/Q9Y287
Images:
https://www.researchgate.net/figure/8910145_fig2_Figure-2-Yeast-protein-interaction-networkA-map-of-protein-protein-interactions18-in
Images:
https://www.researchgate.net/figure/8910145_fig2_Figure-2-Yeast-protein-interaction-networkA-map-of-protein-protein-interactions18-in